

The {fwildclusterboot} package implements multiple fast
wild cluster bootstrap algorithms as developed in Roodman et
al (2019) and MacKinnon,
Nielsen & Webb (2022).
Via the JuliaConnectoR,
{fwildclusterboot} further ports functionality of WildBootTests.jl
- which provides an even faster implementation of the wild cluster
bootstrap for OLS and supports the WRE bootstrap for IV and tests of
multiple joint hypotheses.
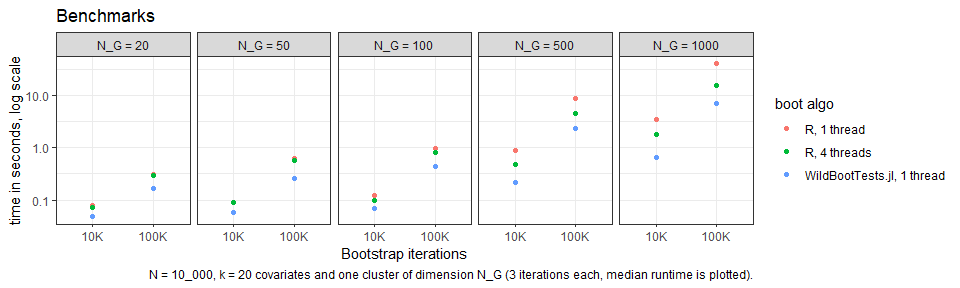
The package’s central function is boottest(). It allows
to test univariate hypotheses using a wild cluster bootstrap at extreme
speed: via the ‘fast’ algorithm, it is possible to run a wild cluster
bootstrap with \(B = 100.000\)
iterations in less than a second!
{fwildclusterboot} supports the following features:
Additional features are provided through
WildBootTests.jl:
{fwildclusterboot} supports the following models:
lm (from stats), fixest (from
fixest), felm from (lfe)ivreg (from ivreg).You can install compiled versions of{fwildclusterboot}
from CRAN (compiled), R-universe (compiled) or github by following one
of the steps below:
# from CRAN
install.packages("fwildclusterboot")
# from r-universe (windows & mac, compiled R > 4.0 required)
install.packages('fwildclusterboot', repos ='https://s3alfisc.r-universe.dev')
# dev version from github
# note: installation requires Rtools
library(devtools)
install_github("s3alfisc/fwildclusterboot")boottest() functionFor a longer introduction to {fwildclusterboot}, take a
look at the vignette.
library(fwildclusterboot)
# set seed via dqset.seed for engine = "R" & Rademacher, Webb & Normal weights
dqrng::dqset.seed(2352342)
# set 'familiar' seed for all other algorithms and weight types
set.seed(23325)
data(voters)
# fit the model via fixest::feols(), lfe::felm() or stats::lm()
lm_fit <- lm(proposition_vote ~ treatment + log_income + as.factor(Q1_immigration) + as.factor(Q2_defense), data = voters)
# bootstrap inference via boottest()
lm_boot <- boottest(lm_fit, clustid = c("group_id1"), B = 9999, param = "treatment")
#> Too guarantee reproducibility, don't forget to set a global random seed
#> **both** via `set.seed()` and `dqrng::dqset.seed()`.
#> This message is displayed once every 8 hours.
summary(lm_boot)
#> boottest.lm(object = lm_fit, param = "treatment", B = 9999, clustid = c("group_id1"))
#>
#> Hypothesis: 1*treatment = 0
#> Observations: 300
#> Bootstr. Type: rademacher
#> Clustering: 1-way
#> Confidence Sets: 95%
#> Number of Clusters: 40
#>
#> term estimate statistic p.value conf.low conf.high
#> 1 1*treatment = 0 0.079 3.983 0.001 0.039 0.119If you are in R, you can simply run the following
command to get the BibTeX citation for
{fwildclusterboot}:
citation("fwildclusterboot")
#>
#> To cite 'fwildclusterboot' in publications use:
#>
#> Fischer & Roodman. (2021). fwildclusterboot: Fast Wild Cluster
#> Bootstrap Inference for Linear Regression Models. Available from
#> https://cran.r-project.org/package=fwildclusterboot.
#>
#> A BibTeX entry for LaTeX users is
#>
#> @Misc{,
#> title = {fwildclusterboot: Fast Wild Cluster Bootstrap Inference for Linear Regression Models (Version 0.12.4.3)},
#> author = {Alexander Fischer and David Roodman},
#> year = {2021},
#> url = {https://cran.r-project.org/package=fwildclusterboot},
#> }Alternatively, if you prefer to cite the “Fast & Wild” paper by
Roodman et al, it would be great if you mentioned
{fwildclusterboot} in a footnote =) !