
pre:
an R package for deriving prediction rule ensemblespre is an R package
for deriving prediction rule ensembles for binary, multinomial,
(multivariate) continuous, count and survival responses. Input variables
may be numeric, ordinal and categorical. An extensive description of the
implementation and functionality is provided in Fokkema (2020). The
package largely implements the algorithm for deriving prediction rule
ensembles as described in Friedman & Popescu (2008), with several
adjustments:
R based,
allowing users better access to the results and more control over the
parameters used for generating the prediction rule ensemble.gpe().Note that pre is under development, and
much work still needs to be done. Below, an introduction the the package
is provided. Fokkema (2020) provides an extensive description of the
fitting procedures implemented in function pre() and
example analyses with more extensive explanations. An extensive
introduction aimed at researchers in social sciences is provided in
Fokkema & Strobl (2020).
To get a first impression of how function pre() works,
we will predict Ozone levels using the airquality dataset.
We fit a prediction rule ensemble using function pre():
library("pre")
airq <- airquality[complete.cases(airquality), ]
set.seed(42)
airq.ens <- pre(Ozone ~ ., data = airq)Note that it is necessary to set the random seed, to allow for later replication of results, because the fitting procedure depends on random sampling of training observations.
We can print the resulting ensemble (alternatively, we could use the
print method):
airq.ens
#>
#> Final ensemble with cv error within 1se of minimum:
#>
#> lambda = 3.543968
#> number of terms = 12
#> mean cv error (se) = 352.395 (99.13754)
#>
#> cv error type : Mean-Squared Error
#>
#> rule coefficient description
#> (Intercept) 68.48270407 1
#> rule191 -10.97368180 Wind > 5.7 & Temp <= 87
#> rule173 -10.90385519 Wind > 5.7 & Temp <= 82
#> rule42 -8.79715538 Wind > 6.3 & Temp <= 84
#> rule204 7.16114781 Wind <= 10.3 & Solar.R > 148
#> rule10 -4.68646145 Temp <= 84 & Temp <= 77
#> rule192 -3.34460038 Wind > 5.7 & Temp <= 87 & Day <= 23
#> rule51 -2.27864287 Wind > 5.7 & Temp <= 84
#> rule93 2.18465676 Temp > 77 & Wind <= 8.6
#> rule74 -1.36479545 Wind > 6.9 & Temp <= 84
#> rule28 -1.15326093 Temp <= 84 & Wind > 7.4
#> rule25 -0.70818400 Wind > 6.3 & Temp <= 82
#> rule166 -0.04751152 Wind > 6.9 & Temp <= 82The first few lines of the printed results provide the penalty
parameter value (λ) employed for selecting the final ensemble.
By default, the ‘1-SE’ rule is used for selecting λ; this
default can be overridden by employing the penalty.par.val
argument of the print method and other functions in the
package. Note that the printed cross-validated error is calculated using
the same data as was used for generating the rules and likely provides
an overly optimistic estimate of future prediction error. To obtain a
more realistic prediction error estimate, we will use function
cvpre() later on.
Next, the printed results provide the rules and linear terms selected
in the final ensemble, with their estimated coefficients. For rules, the
description column provides the conditions. For linear
terms (which were not selected in the current ensemble), the winsorizing
points used to reduce the influence of outliers on the estimated
coefficient would be printed in the description column. The
coefficient column presents the estimated coefficient.
These are regression coefficients, reflecting the expected increase in
the response for a unit increase in the predictor, keeping all other
predictors constant. For rules, the coefficient thus reflects the
difference in the expected value of the response when the conditions of
the rule are met, compared to when they are not.
Using the plot method, we can plot the rules in the
ensemble as simple decision trees. Here, we will request the nine most
important baselearners through specification of the nterms
argument. Through the cex argument, we specify the size of
the node and path labels:
plot(airq.ens, nterms = 9, cex = .5)
Using the coef method, we can obtain the estimated
coefficients for each of the baselearners (we only print the first six
terms here for space considerations):
coefs <- coef(airq.ens)
coefs[1:6, ]
#> rule coefficient description
#> 201 (Intercept) 68.482704 1
#> 167 rule191 -10.973682 Wind > 5.7 & Temp <= 87
#> 150 rule173 -10.903855 Wind > 5.7 & Temp <= 82
#> 39 rule42 -8.797155 Wind > 6.3 & Temp <= 84
#> 179 rule204 7.161148 Wind <= 10.3 & Solar.R > 148
#> 10 rule10 -4.686461 Temp <= 84 & Temp <= 77We can generate predictions for new observations using the
predict method (only the first six predicted values are
printed here for space considerations):
predict(airq.ens, newdata = airq[1:6, ])
#> 1 2 3 4 7 8
#> 32.53896 24.22456 24.22456 24.22456 31.38570 24.22456Using function cvpre(), we can assess the expected
prediction error of the fitted PRE through k-fold cross
validation (k = 10, by default, which can be overridden through
specification of the k argument):
set.seed(43)
airq.cv <- cvpre(airq.ens)
#> $MSE
#> MSE se
#> 369.92747 88.65343
#>
#> $MAE
#> MAE se
#> 13.666860 1.290329The results provide the mean squared error (MSE) and mean absolute
error (MAE) with their respective standard errors. These results are
saved for later use in aiq.cv$accuracy. The cross-validated
predictions, which can be used to compute alternative estimates of
predictive accuracy, are saved in airq.cv$cvpreds. The
folds to which observations were assigned are saved in
airq.cv$fold_indicators.
For tuning the parameters of function pre() so as to
obtain optimal predictive accuracy, users are advised to use
R package
caret. A tutorial is provided as a
vignette, accessible by typing
vignette("tuning", package = "pre") in
R or by going to https://cran.r-project.org/package=pre/vignettes/missingness.html
in a browser and clicking on the corresponding link to the vignette.
Package pre provides several additional
tools for interpretation of the final ensemble. These may be especially
helpful for complex ensembles containing many rules and linear
terms.
We can assess the relative importance of input variables as well as
baselearners using the importance() function:
imps <- importance(airq.ens, round = 4)
As we already observed in the printed ensemble, the plotted variable
importances indicate that Temperature and Wind are most strongly
associated with Ozone levels. Solar.R and Day are also associated with
Ozone levels, but much less strongly. Variable Month is not plotted,
which means it obtained an importance of zero, indicating that it is not
associated with Ozone levels. We already observed this in the printed
ensemble: Month did not appear in any of the selected terms. The
variable and baselearner importances are saved for later use in
imps$varimps and imps$baseimps,
respectively.
We can obtain explanations of the predictions for individual
observations using function explain():
par(mfrow = c(1, 2))
expl <- explain(airq.ens, newdata = airq[1:2, ], cex = .8)
The values of the rules and linear terms for each observation are
saved in expl$predictors, their contributions in
expl$contribution and the predicted values in
expl$predicted.value.
We can obtain partial dependence plots to assess the effect of single
predictor variables on the outcome using the singleplot()
function:
singleplot(airq.ens, varname = "Temp")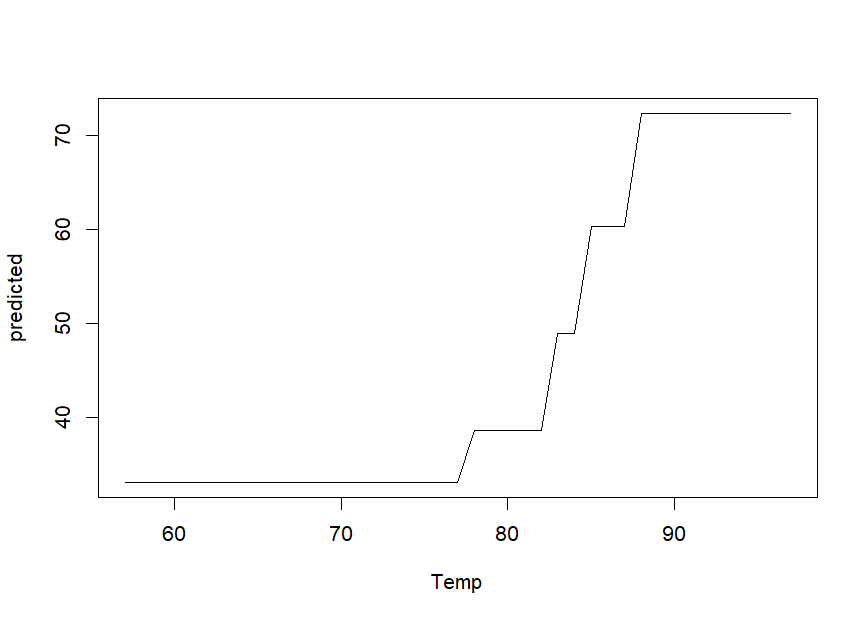
We can obtain partial dependence plots to assess the effects of pairs
of predictor variables on the outcome using the pairplot()
function:
pairplot(airq.ens, varnames = c("Temp", "Wind"))
Note that creating partial dependence plots is computationally
intensive and computation time will increase fast with increasing
numbers of observations and numbers of variables. Milborrow’s (2018)
plotmo package provides more efficient
functions for plotting partial dependence, which also support
pre models.
If the final ensemble does not contain many terms, inspecting
individual rules and linear terms through the print method
may be more informative than partial dependence plots. One of the main
advantages of prediction rule ensembles is their interpretability: the
predictive model contains only simple functions of the predictor
variables (rules and linear terms), which are easy to grasp. Partial
dependence plots are often much more useful for interpretation of
complex models, like random forests for example.
We can assess the presence of interactions between the input
variables using the interact() and
bsnullinteract() funtions. Function
bsnullinteract() computes null-interaction models (10, by
default) based on bootstrap-sampled and permuted datasets. Function
interact() computes interaction test statistics for each
predictor variables appearing in the specified ensemble. If
null-interaction models are provided through the nullmods
argument, interaction test statistics will also be computed for the
null-interaction model, providing a reference null distribution.
Note that computing null interaction models and interaction test statistics is computationally very intensive, so running the following code will take some time:
set.seed(44)
nullmods <- bsnullinteract(airq.ens)
int <- interact(airq.ens, nullmods = nullmods)
The plotted variable interaction strengths indicate that Temperature
and Wind may be involved in interactions, as their observed interaction
strengths (darker grey) exceed the upper limit of the 90% confidence
interval (CI) of interaction stengths in the null interaction models
(lighter grey bar represents the median, error bars represent the 90%
CIs). The plot indicates that Solar.R and Day are not involved in any
interactions. Note that computation of null interaction models is
computationally intensive. A more reliable result can be obtained by
computing a larger number of boostrapped null interaction datasets, by
setting the nsamp argument of function
bsnullinteract() to a larger value (e.g., 100).
We can assess correlations between the baselearners appearing in the
ensemble using the corplot() function:
corplot(airq.ens)
To obtain an optimal set of model-fitting parameters, package
caret Kuhn (2008) provides a method
"pre". For a manual on how to optimize the parameters of
function pre using caret’s
train function, see the vignette on tuning:
vignette("tuning", package = "pre")or go to https://cran.r-project.org/package=pre/vignettes/tuning.html in a browser.
Some suggestions on how to deal with missing values are provided in the following vignette:
vignette("missingness", package = "pre")or go to https://cran.r-project.org/package=pre/vignettes/missingness.html in a browser.
When sparsity (i.e., a final ensemble comprising only few terms) is of central importance, the so-called relaxed lasso can be employed. It allows for retaining a pre-specified (low) number of terms, with adequate predictive accuracy. An introduction and tutorial is provided in the following vignette:
vignette("relaxed", package = "pre")An even more flexible ensembling approach is implemented in function
gpe(), which allows for fitting Generalized Prediction
Ensembles: It combines the MARS (multivariate Adaptive Splines) approach
of Friedman (1991) with the RuleFit approach of Friedman & Popescu
(2008). In other words, gpe() fits an ensemble composed of
hinge functions (possibly multivariate), prediction rules and linear
functions of the predictor variables. See the following example:
set.seed(42)
airq.gpe <- gpe(Ozone ~ ., data = airquality[complete.cases(airquality),],
base_learners = list(gpe_trees(), gpe_linear(), gpe_earth()))
airq.gpe
#>
#> Final ensemble with cv error within 1se of minimum:
#>
#> lambda = 3.229132
#> number of terms = 11
#> mean cv error (se) = 359.2623 (110.8863)
#>
#> cv error type : Mean-squared Error
#>
#> description coefficient
#> (Intercept) 65.52169488
#> Temp <= 77 -6.20973855
#> Wind <= 10.3 & Solar.R > 148 5.46410966
#> Wind > 5.7 & Temp <= 82 -8.06127415
#> Wind > 5.7 & Temp <= 84 -7.16921733
#> Wind > 5.7 & Temp <= 87 -8.04255471
#> Wind > 5.7 & Temp <= 87 & Day <= 23 -3.40525575
#> Wind > 6.3 & Temp <= 82 -2.71925536
#> Wind > 6.3 & Temp <= 84 -5.99085126
#> Wind > 6.9 & Temp <= 82 -0.04406376
#> Wind > 6.9 & Temp <= 84 -0.55827336
#> eTerm(Solar.R * h(9.7 - Wind), scale = 410) 9.91783320
#>
#> 'h' in the 'eTerm' indicates the hinge functionThe results indicate that several rules, a single hinge (linear spline) function, and no linear terms were selected for the final ensemble. The hinge function and its coefficient indicate that Ozone levels increase with increasing solar radiation and decreasing wind speeds. The prediction rules in the ensemble indicate a similar pattern.
I am very grateful to package co-author Benjamin Chistoffersen: https://github.com/boennecd, who developed
gpe and contributed tremendously by improving functions,
code and computational aspects. Furthermore, I am grateful for the many
helpful suggestions of Stephen Milborrow, and for the contributions of
Karl Holub (https://github.com/holub008) and Advik Shreekumar (https://github.com/adviksh).